mutate() adds new variables and preserves existing ones.
select() keeps only the listed variables; rename() keeps all variables.
Arguments
- .data
An xpose database object.
- ...
Name-value pairs of expressions. Use
NULLto drop a variable.These arguments are automatically quoted and evaluated in the context of the data frame. They support unquoting and splicing. See the dplyr vignette("programming") for an introduction to these concepts.
- .problem
The problem from which the data will be modified
- .source
The source of the data in the xpdb. Can either be 'data' or an output file extension e.g. 'phi'.
- .where
A vector of element names to be edited in special (e.g.
.where = c('vpc_dat', 'aggr_obs')with vpc).
Examples
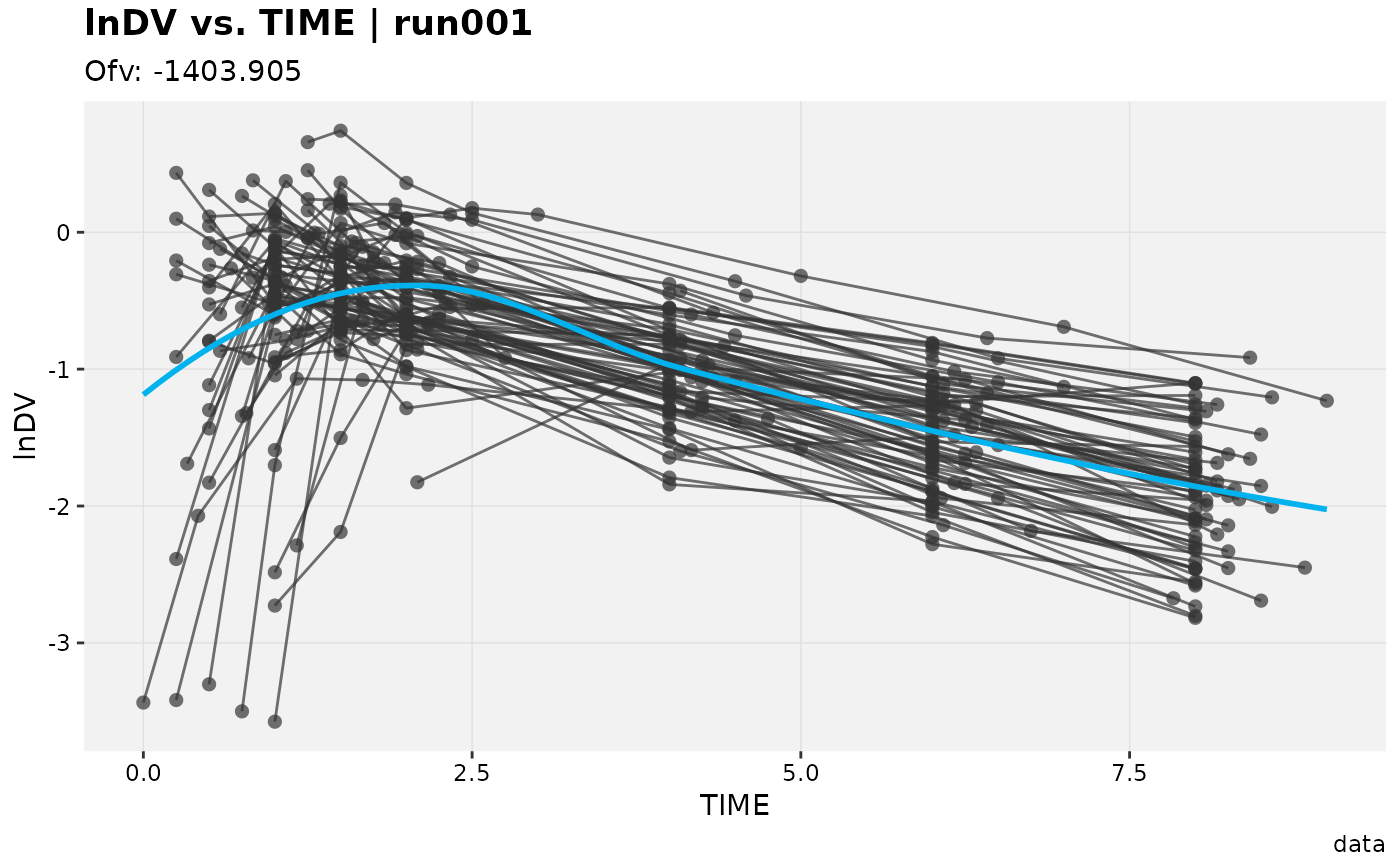
# Mutate columns
xpdb_ex_pk %>%
mutate(lnDV = log(DV),
sim_count = irep(ID),
.problem = 1) %>%
dv_vs_idv(aes(y = lnDV))
#> irep: 1 simulations found.
#> Using data from $prob no.1
#> Filtering data by EVID == 0
#> `geom_smooth()` using formula = 'y ~ x'
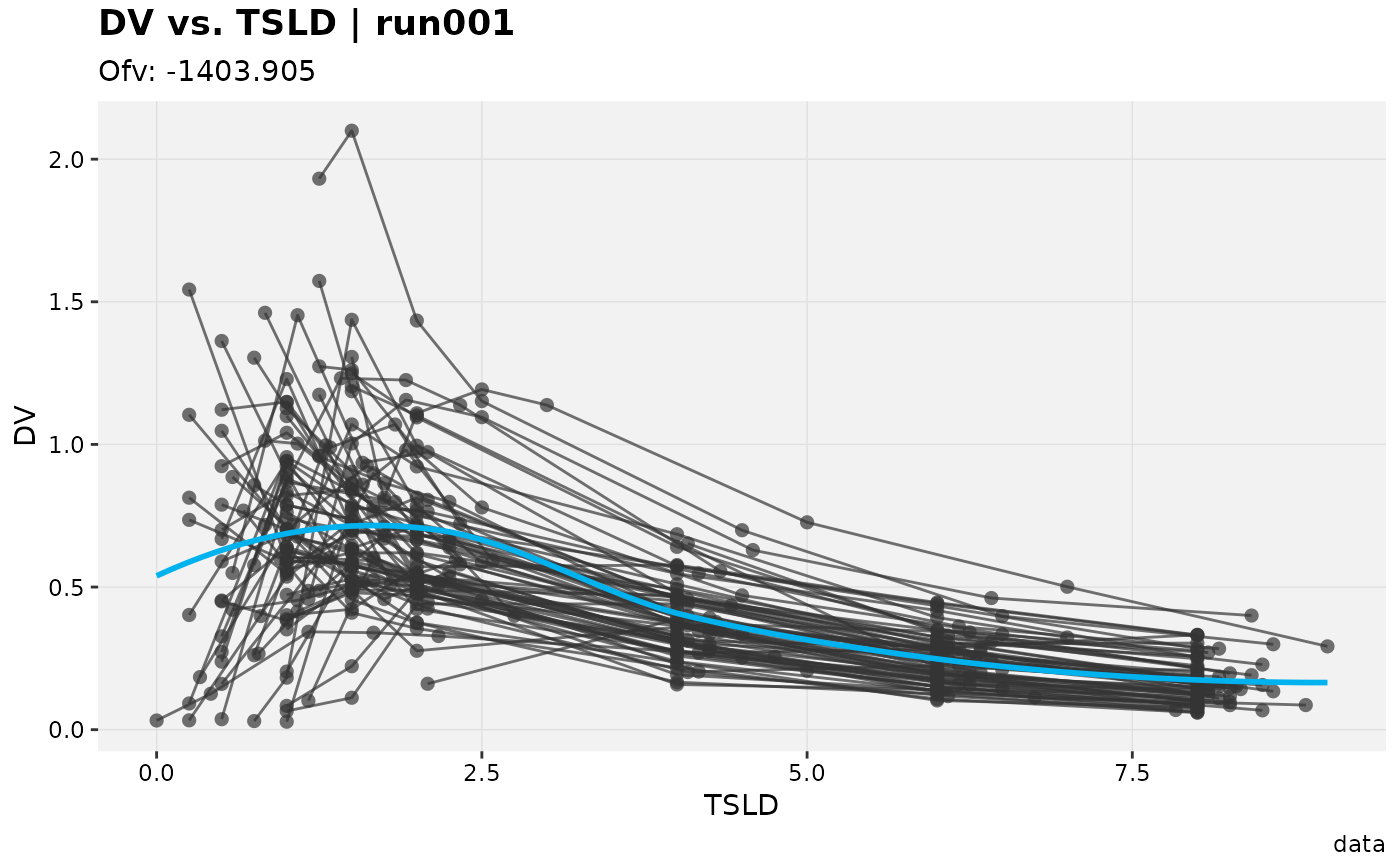
 # Rename/select columns
xpdb_ex_pk %>%
select(ID:TAD, DV, EVID) %>%
rename(TSLD = TAD) %>%
dv_vs_idv(aes(x = TSLD))
#> Using data from $prob no.1
#> Filtering data by EVID == 0
#> `geom_smooth()` using formula = 'y ~ x'
# Rename/select columns
xpdb_ex_pk %>%
select(ID:TAD, DV, EVID) %>%
rename(TSLD = TAD) %>%
dv_vs_idv(aes(x = TSLD))
#> Using data from $prob no.1
#> Filtering data by EVID == 0
#> `geom_smooth()` using formula = 'y ~ x'